|
ATCC
ln-18 Ln 18, supplied by ATCC, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ln-18/product/ATCC Average 96 stars, based on 1 article reviews
ln-18 - by Bioz Stars,
2026-04
96/100 stars
|
Buy from Supplier |
|
Novus Biologicals
pad2 Pad2, supplied by Novus Biologicals, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pad2/product/Novus Biologicals Average 90 stars, based on 1 article reviews
pad2 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
ModiQuest
recombinant human pad2  Recombinant Human Pad2, supplied by ModiQuest, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/recombinant human pad2/product/ModiQuest Average 90 stars, based on 1 article reviews
recombinant human pad2 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Proteintech
anti padi2  Anti Padi2, supplied by Proteintech, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/anti padi2/product/Proteintech Average 94 stars, based on 1 article reviews
anti padi2 - by Bioz Stars,
2026-04
94/100 stars
|
Buy from Supplier |
|
Cayman Chemical
recombinant human pad2  Recombinant Human Pad2, supplied by Cayman Chemical, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/recombinant human pad2/product/Cayman Chemical Average 90 stars, based on 1 article reviews
recombinant human pad2 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Proteintech
pad2  Pad2, supplied by Proteintech, used in various techniques. Bioz Stars score: 95/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pad2/product/Proteintech Average 95 stars, based on 1 article reviews
pad2 - by Bioz Stars,
2026-04
95/100 stars
|
Buy from Supplier |
|
Proteintech
anti pad2  Anti Pad2, supplied by Proteintech, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/anti pad2/product/Proteintech Average 94 stars, based on 1 article reviews
anti pad2 - by Bioz Stars,
2026-04
94/100 stars
|
Buy from Supplier |
|
ProteoGenix
murine pad2 pad4  Murine Pad2 Pad4, supplied by ProteoGenix, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/murine pad2 pad4/product/ProteoGenix Average 90 stars, based on 1 article reviews
murine pad2 pad4 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Millipore
rabbit skeletal peptidylarginine deiminase-2 (pad2  Rabbit Skeletal Peptidylarginine Deiminase 2 (Pad2, supplied by Millipore, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/rabbit skeletal peptidylarginine deiminase-2 (pad2/product/Millipore Average 90 stars, based on 1 article reviews
rabbit skeletal peptidylarginine deiminase-2 (pad2 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Talecris Biotherapeutics
human serum albumin hsa  Human Serum Albumin Hsa, supplied by Talecris Biotherapeutics, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/human serum albumin hsa/product/Talecris Biotherapeutics Average 90 stars, based on 1 article reviews
human serum albumin hsa - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Merck & Co
f95 pan-deimination antibody  Table S1 for full details on all protein hits)." width="250" height="auto" /> Table S1 for full details on all protein hits)." width="250" height="auto" />F95 Pan Deimination Antibody, supplied by Merck & Co, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/f95 pan-deimination antibody/product/Merck & Co Average 90 stars, based on 1 article reviews
f95 pan-deimination antibody - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
ModiQuest
recombinant human pad4  Table S1 for full details on all protein hits)." width="250" height="auto" /> Table S1 for full details on all protein hits)." width="250" height="auto" />Recombinant Human Pad4, supplied by ModiQuest, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/recombinant human pad4/product/ModiQuest Average 90 stars, based on 1 article reviews
recombinant human pad4 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: Journal of immunology (Baltimore, Md. : 1950)
Article Title: Citrullination alters immunomodulatory function of LL-37 essential for prevention of endotoxin-induced sepsis
doi: 10.4049/jimmunol.1303062
Figure Lengend Snippet: Human MDMs (A) and mouse RAW 264.7 (B) macrophages were stimulated with 10 ng/mL LPS in the presence of native or citrullinated LL-37 at the indicated concentrations (0.1–10 μg/mL). Citrullinated LL-37 was obtained by treatment of the native peptide with human PAD2 or PAD4 at 23.3 U/mg peptide). The level of TNF-α (A) and NO (B) in the culture supernatants was determined using ELISA or the Griess assay at 6 or 20 h post-stimulation, respectively. Since neither the LL-37 nor the PAD enzymes alone induced the release of NO or TNF-α, for the sake of clarity these controls are not shown in the figure. Data represent the mean ± SD of three independent experiments. ns, not significant; *, p<0.05; **, p<0.01; ***, p<0.001.
Article Snippet:
Techniques: Enzyme-linked Immunosorbent Assay, Griess Assay

Journal: bioRxiv
Article Title: Citrullination of TDP-43 is a key post-translation modification associated with structural and functional changes and progressive pathology in TDP-43 mouse models and human proteinopathies
doi: 10.1101/2025.02.28.639952
Figure Lengend Snippet: a, The enzymatic reaction mediated by PADs converts arginine to citrulline. b, c, Western blot images of PAD2 and PAD4-mediated citrullination of H3 (b) and TDP-43 (c). In (c) arrows indicate the observed shift in TDP-43 molecular weight. d, Coomassie staining of PAD2 (bands 1, 4), PAD4 (bands 2, 6), unmodified TDP-43 (band 3), PAD2-mediated citR TDP-43 (band 5), and PAD4-mediated citR TDP-43 (band 7). e, Bar diagrams showing the MW shift (kDa) of TDP-43 protein following citrullination. f, Schematic representation of arginine epitopes positioned in the human TDP-43 protein sequence. Position of the 11 citrullinated arginine epitopes are indicated in red. g, Six out of eleven arginine epitopes susceptible to citrullination laid within common RX(X)R (red box) or RXG/RGGG (green box) motifs in TDP-43 sequence. h, MS/MS spectrum showing b- and y-ion coverage of modified citR83 peptide and extracted ion chromatograms (XICs) showed abundance and retention time of unmodified R83 vs. citR83 with PAD2 and PAD4 treatment (intact peptide monoisotopic m/z (+2) 719.8457, (+3) 480.2329). i, Retention time peaks for TDP-43 peptides surrounding the unmodified or modified R83 epitope and base peak m/z undergoing methionine (Met85) oxidation, citrullination or both showed reliable time separation.
Article Snippet:
Techniques: Western Blot, Molecular Weight, Staining, Sequencing, Tandem Mass Spectroscopy, Modification

Journal: bioRxiv
Article Title: Citrullination of TDP-43 is a key post-translation modification associated with structural and functional changes and progressive pathology in TDP-43 mouse models and human proteinopathies
doi: 10.1101/2025.02.28.639952
Figure Lengend Snippet: Spectra showing b- and y-ion coverage of citrullinated peptides, as well as XICs retention times for unmodified and citrullinated peptides treated with PAD2 and PAD4, corresponding to a - c, citR165 (intact peptide monoisotopic m/z (+2) 665.8214), d - f, citR191(intact peptide monoisotopic m/z (+2) 672.8469), g - i, citR268/272 (intact peptide monoisotopic m/z (+2) 728.3560), j - l, citR293 (intact peptide monoisotopic m/z (+2) 638.29).
Article Snippet:
Techniques:
Journal: bioRxiv
Article Title: Citrullination of TDP-43 is a key post-translation modification associated with structural and functional changes and progressive pathology in TDP-43 mouse models and human proteinopathies
doi: 10.1101/2025.02.28.639952
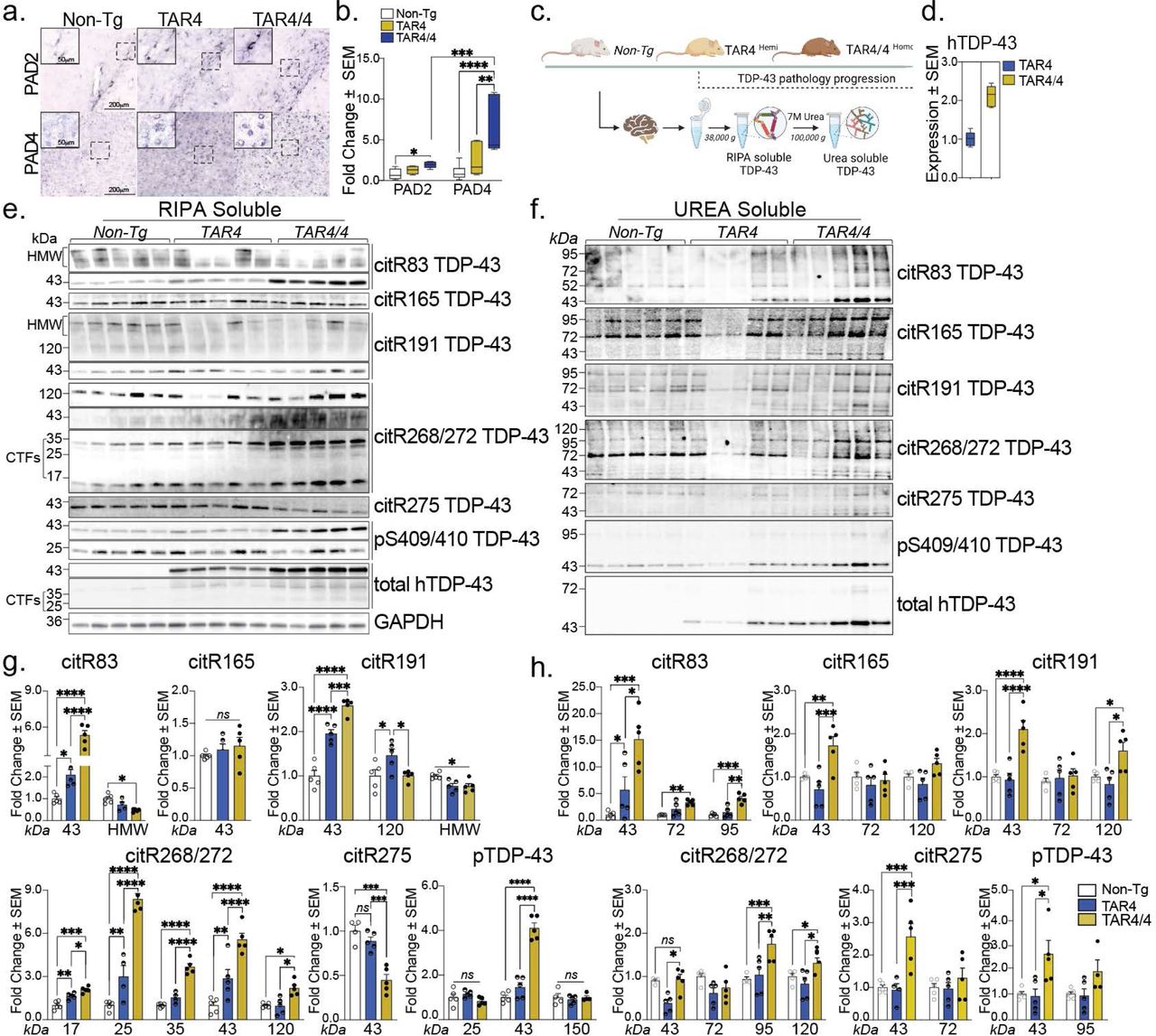
Figure Lengend Snippet: a, Immunohistochemical images of Non-Tg, TAR4 and TAR4/4 cortex labeled with PAD2 and PAD4 antibodies. b, Fold change of PAD2 and PAD4 (% area mean of the values ± SEM) normalized to the Non-Tg control. One-way or Two-way ANOVA, followed by Tukey’s post hoc multiple comparisons tests, n = 5, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. c, Schematic overview of the TAR TDP-43 mouse model and the fractionation of cortical tissue into RIPA-soluble and (7 M) Urea-soluble fractions. d, The expression levels of the human TDP-43 in the TAR model. e, Cortical RIPA soluble and f, Urea soluble fraction analyzed by Western blotting and probed for citR TDP-43 antibody panel (citR83, 165, 191, 268/272, 275), pTDP-43 409/410 and total human TDP-43 protein. g, Quantification of citR TDP-43 43 kDa protein levels, proteolytic fragments (17, 25 & 35 kDa) and intermediate high molecular species (72, 98 & 120k Da) normalized to GAPDH in RIPA and h, Urea fraction normalized to the total protein loaded signal. Data represent the mean of the values ± SEM; One-way ANOVA, followed by Tukey’s or Šidák post hoc multiple comparisons tests, n = 5, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Article Snippet:
Techniques: Immunohistochemical staining, Labeling, Control, Fractionation, Expressing, Western Blot

Journal: Proceedings of the National Academy of Sciences of the United States of America
Article Title: Peptidyl arginine deiminase immunization induces anticitrullinated protein antibodies in mice with particular MHC types
doi: 10.1073/pnas.1713112114
Figure Lengend Snippet: IgG responses to PADs in C3H mice. IgG responses to murine PAD2 or human PAD2 or human PAD4 were analyzed by ELISAs. Plates were coated with PADs and blocked with BSA. Sera from primed mice were obtained at 15, 35, 55, and 65 d postimmunization and were diluted at 1/40. ( A ) For PBS-immunized mice, 15 sera were tested for murine PAD2, 8 sera were tested for human PAD2, and 9 sera were tested for human PAD4. ( B ) For PAD-immunized mice, each serum was tested against the same PAD used for each immunization. After washing, peroxidase-conjugated antimurine IgG was added. The OD was read at 405 nm. The background OD was obtained by adding each serum to a well without PAD (negative). Positive sera were defined as an OD value higher than twice the background OD.
Article Snippet:
Techniques:

Journal: Proceedings of the National Academy of Sciences of the United States of America
Article Title: Peptidyl arginine deiminase immunization induces anticitrullinated protein antibodies in mice with particular MHC types
doi: 10.1073/pnas.1713112114
Figure Lengend Snippet: IgG responses to PADs in DBA/2 mice. Plates were coated with PADs and blocked with BSA. Sera from primed mice were obtained at 15, 35, 55, and 65 d postimmunization and were diluted at 1/40. ( A ) For PBS-immunized mice, nine sera were tested for murine PAD2 and human PAD4. ( B ) For PAD-immunized mice, each serum was tested against the same PAD used for each immunization. After washing, peroxidase-conjugated antimurine IgG was added. The OD was read at 405 nm. The background OD was obtained by adding each serum to a well without PAD (negative). Positive sera were defined as an OD value higher than twice the background OD.
Article Snippet:
Techniques:
 Table S1 for full details on all protein hits)." width="100%" height="100%">
Table S1 for full details on all protein hits)." width="100%" height="100%">
Journal: Biology
Article Title: Extracellular Vesicles and Post-Translational Protein Deimination Signatures in Mollusca—The Blue Mussel ( Mytilus edulis ), Soft Shell Clam ( Mya arenaria ), Eastern Oyster ( Crassostrea virginica ) and Atlantic Jacknife Clam ( Ensis leei )
doi: 10.3390/biology9120416
Figure Lengend Snippet: Deiminated proteins in hemolymph of blue mussel ( Mytilus edulis ), as identified by F95-enrichment in conjunction with LC–MS/MS analysis. Deiminated proteins were isolated from hemolymph (a pool of hemolymph from four individual animals) by immunoprecipitation using the pan-deimination F95 antibody. The resulting F95-enriched eluate was then analyzed by LC–MS/MS and peak list files submitted to Mascot, using both a species-specific as well as a common Mollusca database. Peptide sequence hits are listed, showing species-specific hits, number of sequences for protein hits, and total score. Species hit names are indicated, blue mussel specific hits are on the top of the list and highlighted. *Proteins only identified in blue mussel. (See
Article Snippet: Primary antibody incubation was carried out overnight at 4 °C on a shaking platform using the following antibodies for Mollusca sera:
Techniques: Isolation, Immunoprecipitation, Sequencing, Binding Assay
 Table S2 for full details on all protein hits)." width="100%" height="100%">
Table S2 for full details on all protein hits)." width="100%" height="100%">
Journal: Biology
Article Title: Extracellular Vesicles and Post-Translational Protein Deimination Signatures in Mollusca—The Blue Mussel ( Mytilus edulis ), Soft Shell Clam ( Mya arenaria ), Eastern Oyster ( Crassostrea virginica ) and Atlantic Jacknife Clam ( Ensis leei )
doi: 10.3390/biology9120416
Figure Lengend Snippet: Deiminated proteins in hemolymph of soft shell clam ( Mya arenaria ), as identified by F95-enrichment in conjunction with LC–MS/MS analysis. Deiminated proteins were isolated from hemolymph (a pool of hemolymph from four individual animals) by immunoprecipitation using the pan-deimination F95 antibody. The resulting F95-enriched eluate was then analyzed by LC–MS/MS and peak list files submitted to Mascot, using both a species-specific as well as a common Mollusca database. Peptide sequence hits are listed, showing species-specific hit, number of sequences for protein hits, and total score. Species hit names are indicated, soft shell clam specific hits are on the top of the list and highlighted. * Proteins only identified in soft shell clam. (See
Article Snippet: Primary antibody incubation was carried out overnight at 4 °C on a shaking platform using the following antibodies for Mollusca sera:
Techniques: Isolation, Immunoprecipitation, Sequencing
 Table S3 for full details on all protein hits)." width="100%" height="100%">
Table S3 for full details on all protein hits)." width="100%" height="100%">
Journal: Biology
Article Title: Extracellular Vesicles and Post-Translational Protein Deimination Signatures in Mollusca—The Blue Mussel ( Mytilus edulis ), Soft Shell Clam ( Mya arenaria ), Eastern Oyster ( Crassostrea virginica ) and Atlantic Jacknife Clam ( Ensis leei )
doi: 10.3390/biology9120416
Figure Lengend Snippet: Deiminated proteins in hemolymph of Eastern oyster ( Crassostrea virginica ), as identified by F95-enrichment followed by LC–MS/MS analysis. Deiminated proteins were isolated from hemolymph (a pool of hemolymph from four individual animals) by immunoprecipitation using the pan-deimination F95 antibody. The resulting F95-enriched eluate was then analyzed by LC–MS/MS and peak list files submitted to Mascot, using both a species-specific as well as a common Mollusca database. Peptide sequence hits are listed, showing species-specific hit, number of sequences for protein hits, and total score. Species hit names are indicated, Eastern oyster specific hits are on the top of the list and highlighted. * Proteins only identified in Eastern oyster. (See
Article Snippet: Primary antibody incubation was carried out overnight at 4 °C on a shaking platform using the following antibodies for Mollusca sera:
Techniques: Isolation, Immunoprecipitation, Sequencing
 Table S4 for full details on all protein hits)." width="100%" height="100%">
Table S4 for full details on all protein hits)." width="100%" height="100%">
Journal: Biology
Article Title: Extracellular Vesicles and Post-Translational Protein Deimination Signatures in Mollusca—The Blue Mussel ( Mytilus edulis ), Soft Shell Clam ( Mya arenaria ), Eastern Oyster ( Crassostrea virginica ) and Atlantic Jacknife Clam ( Ensis leei )
doi: 10.3390/biology9120416
Figure Lengend Snippet: Deiminated proteins in hemolymph of Atlantic jacknife clam ( Ensis leei ), as identified by F95-enrichment followed by LC–MS/MS analysis. Deiminated proteins were isolated from hemolymph (a pool of hemolymph from four individual animals) by immunoprecipitation using the pan-deimination F95 antibody. The resulting F95-enriched eluate was then analyzed by LC–MS/MS and peak list files submitted to Mascot, using both a species-specific as well as a common Mollusca database. Peptide sequence hits are listed, showing species-specific hit, number of sequences for protein hits, and total score. * Proteins only identified in Atlantic jacknife clam. (See
Article Snippet: Primary antibody incubation was carried out overnight at 4 °C on a shaking platform using the following antibodies for Mollusca sera:
Techniques: Isolation, Immunoprecipitation, Sequencing